Workflow Description¶
Workflow definition diagrams¶
The following diagram shows the rules executed by the workflow and their dependencies. Each rule is associated with the execution of either a python script, a jupyter notebook or a bash script.
The actual execution of the workflow requires some of the rules to be executed multiple times. In particular each subcube is processed sequentially. The next diagram shows the DAG of an example execution. The number of parallel jobs is variable, here we show the case of 9 subcubes, although for the full SDC2 cube we may use 36 or 49 subcubes.
Each rule has associated input and output files. The following diagram shows the stage at which the relevant files are created or used.
Workflow file structure¶
The workflow consists of a master Snakefile file (analogous to a Makefile for make), a series of conda environments, scripts and notebooks to be executed, and rules to manage the workflow tasks. The file organization of the workflow is the following:
workflow/
├── Snakefile
├── envs
│ ├── analysis.yml
│ ├── chunk_data.yml
│ ├── filter_catalog.yml
│ ├── process_data.yml
│ ├── snakemake.yml
│ └── xmatch_catalogs.yml
├── notebooks
│ └── sdc2_hi-friends.ipynb
├── rules
│ ├── chunk_data.smk
│ ├── concatenate_catalogs.smk
│ ├── run_sofia.smk
│ ├── summary.smk
│ └── visualize_products.smk
├── scripts
│ ├── define_chunks.py
│ ├── eliminate_duplicates.py
│ ├── filter_catalog.py
│ ├── run_sofia.py
│ ├── sofia2cat.py
│ └── split_subcube.py
Output products¶
All the outputs of the workflow are stored in results. This is the first level organization of the directories:
results/
├── catalogs
├── logs
├── notebooks
├── plots
└── sofia
In particular, each rule generates a log for the execution of the scripts. They are stored in results/logs. Each subdirectory contains individual logs for each executtion, as shown in this example:
logs/
├── concatenate
│ ├── concatenate_catalogs.log
│ ├── eliminate_duplicates.log
│ └── filter_catalog.log
├── define_chunks
│ └── define_chunks.log
├── run_sofia
│ ├── subcube_0.log
│ ├── subcube_1.log
│ ├── subcube_2.log
│ └── subcube_3.log
├── sofia2cat
│ ├── subcube_0.log
│ ├── subcube_1.log
│ ├── subcube_2.log
│ └── subcube_3.log
├── split_subcube
│ ├── subcube_0.log
│ ├── subcube_1.log
│ ├── subcube_2.log
│ └── subcube_3.log
└── visualize
└── visualize.log
The individual fits files of the subcubes are stored in the directory interim because they may be large to store. The workflow can be setup to make them temporary files, so they are removed as soon as they are not needed anymore. After an execution, it is possible to maintain all the relevant outputs by keeping the results directory, while the interim directory, only containing fits files, can be safely removed if not explicitly needed.
interim/
└── subcubes
├── subcube_0.fits
├── subcube_1.fits
├── subcube_2.fits
└── subcube_3.fits
...
Snakemake execution and diagrams¶
Additional files summarizing the execution of the workflow and the Snakemake rules are stored in summary. These are not generated by the main snakemake job, but need to be generated once the main job is finished by executing snakemake specifically for this purpose. The four commands to produce these additional plots are executed by the main script run.py.
summary/
├── dag.svg
├── filegraph.svg
├── report.html
└── rulegraph.svg
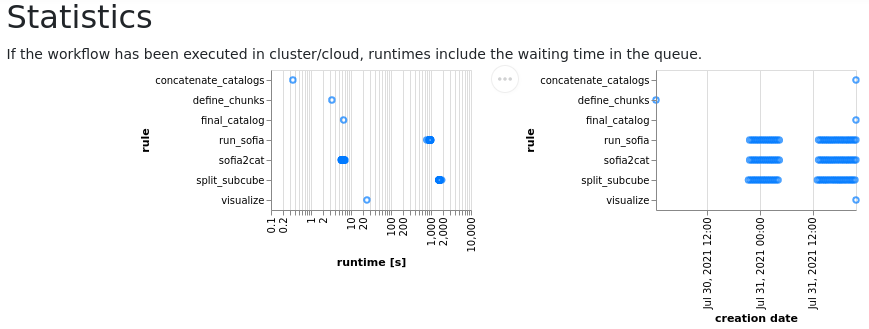
In particular, report.hml contains a description of the rules, including the provenance of each execution, as well as the statistics on execution times of each rule.
Interactive report showing the workflow structure:
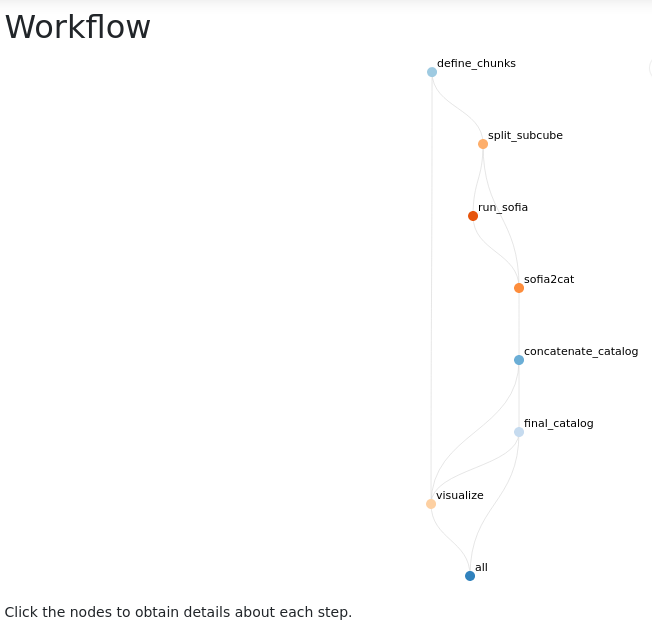
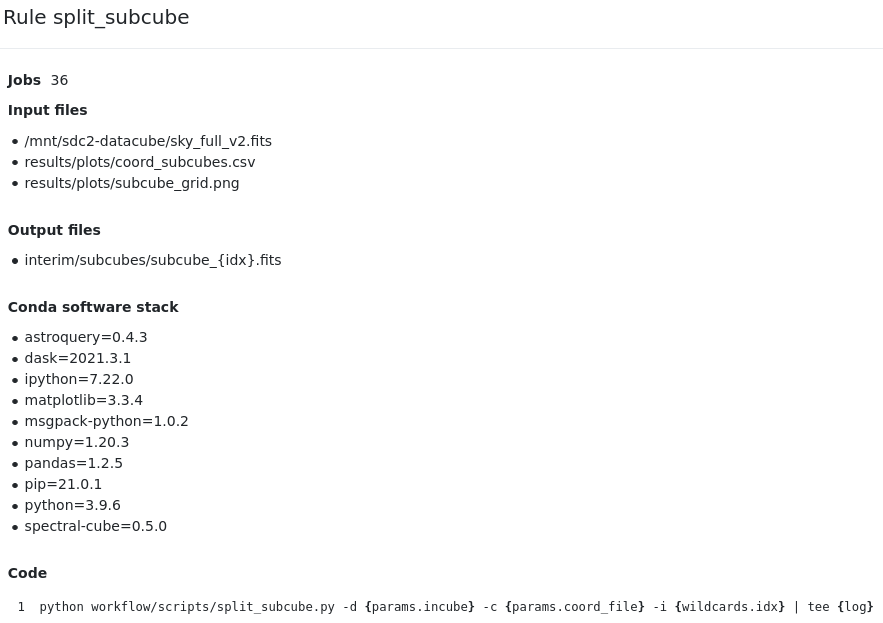
When clicking in one of the nodes, full provenance is provided:
Statistics of the time required for each execution: